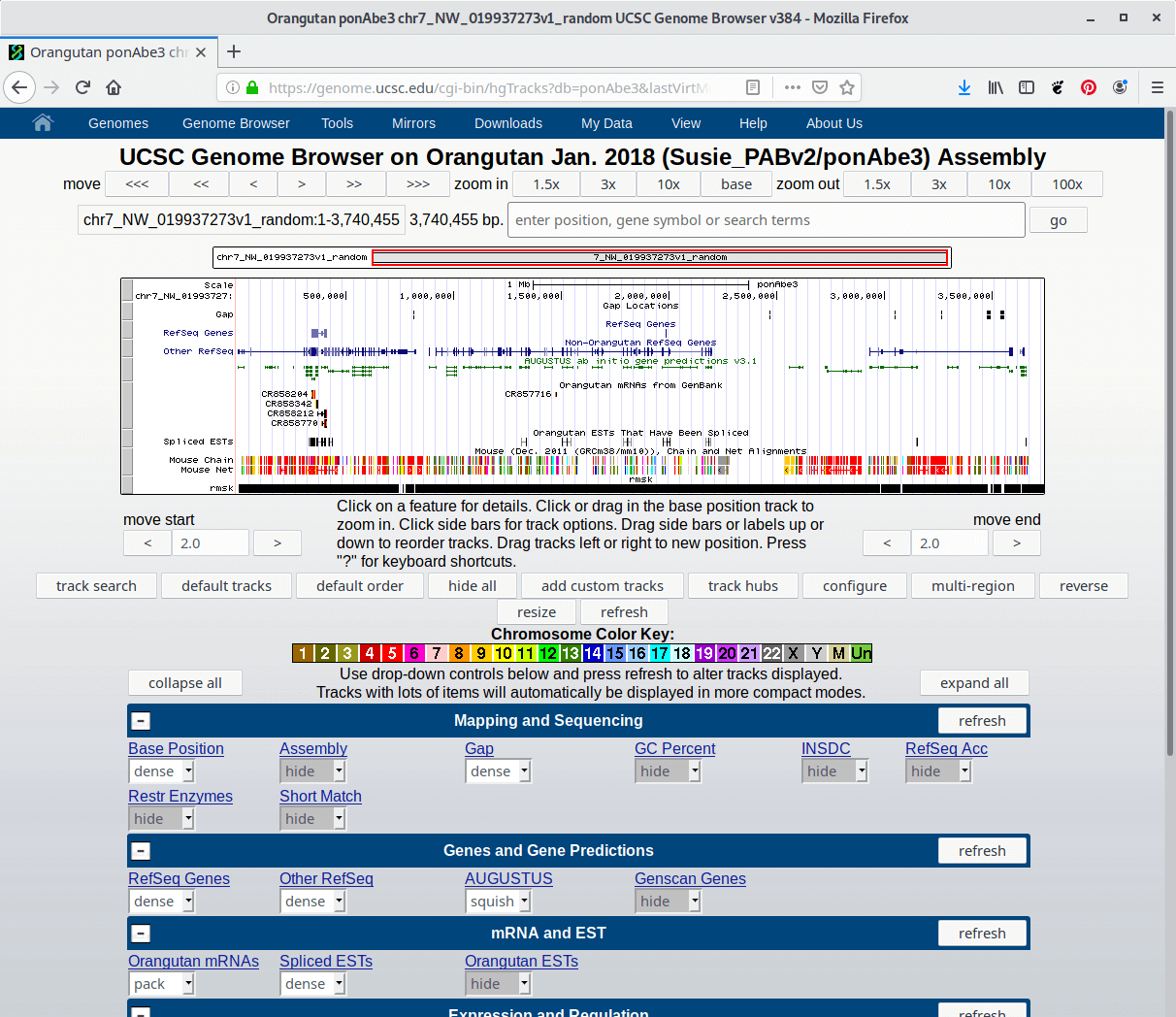
This feature is displayed (higher numbers = darker gray). Score value will determine the level of gray in which UseScore attribute is set to 1 for this annotation data set, the Window when the track is open to full display mode or directly to the This label isĭisplayed to the left of the BED line in the Genome Browser name - Defines the name of the BED line.The 9 additional optional BED fields are: For example, the first 100 bases of aĬhromosome are defined as chromStart=0, chromEnd=100, and span The chromEnd base is not included in theĭisplay of the feature. chromEnd - The ending position of the feature in theĬhromosome or scaffold.The first base in a chromosome is numbered 0. chromStart - The starting position of the feature in theĬhromosome or scaffold.chr3, chrY,Ĭhr2_random) or scaffold (e.g. chrom - The name of the chromosome (e.g.It on your own server, you should use the If your data set is BED-like, but it is very large and you would like to keep Optional fields is binding: lower-numbered fields must always be populated if Throughout any single set of data in an annotation track. The number of fields per line must be consistent 
0 Comments
Leave a Reply. |
 RSS Feed
RSS Feed
